提高玉米(Zea mays L.)耐密性是提高玉米单产的关键措施之一。单位面积玉米种植密度增加往往会导致玉米茎秆变细和倒伏率增加,从而导致产量降低[1-3]。种植密度和茎粗是影响玉米茎秆倒伏的主要因素之一;在高密度下,增加茎粗会降低茎秆倒伏的概率[4]。王铁固等[5]基于2个玉米组合发现作为数量性状的茎粗主要受到1对加性主基因+加性-显性多基因模型或2对加性-显性-上位性主基因+加性-显性多基因混合遗传模型的控制。目前,已有相关研究基于双亲的F2∶3[6]、重组自交系(RIL)[7]、BC6S6[8]、双单倍体[9]、四交群体F2∶3家系[10]和关联群体[11]等遗传群体对玉米茎粗进行遗传解析。徐蔚[6]利用来源掖478×R08的玉米F2∶3群体在四川省雅安市鉴定到1个2号染色体上的控制茎粗的主效数量性状位点(QTL)。Hu等[7]以衍生于玉米组合B73×Ceo3005的RIL为试验材料报道了一个位于bin5.03上同时控制宽和窄茎直径的主效QTL。Mazaheri等[11]使用1个扩展版玉米关联群体(942份自交系)进行茎粗全基因组关联分析检测到了8个与茎粗相关的候选基因,其中2个与花期相关候选基因(Zmm22和Fpa同源)的同源基因还影响拟南芥莲座半径。刘鹏飞[10]在高低密度下对1个玉米四交群体的茎粗及其系数进行QTL分析,发掘到2个影响茎粗系数的位点(bin3.01-3.02和bin10.03-10.04),且后者增效基因来源于耐密型亲本郑58和PH6WC。此外,刘福鹏等[12]通过元分析方法得到20个与玉米茎粗相关的一致性QTL。尽管诸多研究对玉米茎粗进行了遗传分析,但当前对茎粗响应密度的遗传基础报道仍较少。
现有关于玉米耐密性的QTL研究多集中于株型[13-19]、产量[20-23]和花期[13-14,24]等相关性状。相较于玉米种植密度较高的我国中部和北部地区,作为我国第三大玉米产区的西南地区数十年来种植密度水平一直在低水平徘徊。特殊的地域环境使生产者偏好种植稀植大穗型玉米品种,且抗病和不耐密的热带种质与PB种质在该区域使用广泛。因此,本研究利用来源于耐密型骨干自交系掖478和稀植型骨干系R08(或08-641,PB种质)的301个RIL在2014,2015年云南省景洪高低密度下测量茎粗,并在联合环境下进行QTL定位、上位性分析、QTL与环境互作和QTL与密度互作,旨在为改良西南地区玉米耐密性和抗倒伏的遗传基础提供帮助。
1 材料和方法
1.1 试验材料
本研究采用的玉米RIL群体是通过来源于R08×掖478组合的单籽粒传法构建成的301个F8家系(RIL群体)[19-20]。玉米骨干亲本掖 478株型紧凑、矮秆和穗位低,是典型的耐密和抗倒伏的自交系[10]。R08则为西南地区骨干自交系,具有配合力高、制种产量高和抗性较好等优点。但R08株高较高,是西南地区典型稀植大穗型自交系。二者在多数株型和产量相关性状上存在显著差异[19-20]。
1.2 田间试验设计和性状调查
本研究将掖478、R08、掖478×R08及301个RIL种植于2014,2015年云南省景洪2种密度下,并测定茎粗。试验采用完全随机区组设计,2个重复,每个小区一行,行长 3 m。低密度为 57 000 株/hm2,高密度为 114 000 株/hm2。环境、播种、行株距及管理细节详见参考文献[19-20]。本试验在RIL群体自然散粉后15 d开始在田间测量茎粗。从每行随机选择5个的植株(除边株外)用电子游标卡尺测量植株地上部第3节窄截面的茎秆直径(mm)[10]。
1.3 田间数据分析
将每个环境每个密度下得到的5个重复数值计算平均值,随后将其用于双亲性状差异显著性和群体表型分布分析。利用SAS 9.4[25]统计学软件中的混合线性模型PROC MIXED程序进行联合方差分析。其中基因型、基因型与环境互作、基因型与密度互作为随机因子,密度与环境和重复互作、密度与环境互作、环境和密度均为固定因子[20]。高低密度下茎粗以及茎粗与株型和产量相关性状[19-20]的皮尔森相关系数是采用 SAS 9.4[25]的 PROC CORR程序对联合环境下均值估算的。2种密度下基于家系均值的遗传力是利用SAS 9.4[25]基于Holland等[26]方法进行估算的。对于高低密度下的茎粗数据,采用如下混合线性模型估计其最佳线性无偏预测值(BLUP):Ymik=μ+Gm+GEmi+Ei+Rk+εmik。其中,Ymik为第i环境下第k个重复基因型m的表型值;μ是平均值;Gm为基因型m的效应值;GEmi表示第i个环境和基因型m的互作效应值;Ei为第i个环境的效应值;Rk为第k个重复的效应值;εmik为残差。上述所有因子均被视为随机因子。每个密度下RIL的BLUP随后被用于QTL定位分析。BLUP分析采用R语言进行计算[27-28]。
1.4 遗传图谱的构建
本研究遗传图谱参照先前的研究[19-20]。即从每个RIL随机选取10颗种子播于育苗盘内。做遮光处理育苗,7 d后剪下其叶片参照Chen等[29]的方法提取其全基因组DNA。基于MaizeSNP 3K芯片完成基因型分型[30]。根据所得到的SNP分型数据剔除杂合率和缺失率大于 20%以及偏分离严重的标记以及杂合率过高的自交系。最后获得了301个RIL和 683个高质量的SNP标记。用MapDisto 1.7.5[31]构建遗传连锁图谱。用Kosambi作图函数转换重组率为遗传距离[32]。总遗传图谱长为1 786.06 cM。
1.5 QTL定位、上位性、QTL与环境及密度互作分析
本研究结合RIL群体基因型和BLUP值并采用基于复合区间作图法(ICIM)的QTL ICIMapping软件[33]进行 QTL定位分析。标记变量残差表型逐步回归中的扫描步长及输入变量的P值分别设定为 1 cM和 0.001。去除变量的最大P值被假定为2倍的PIN值。阈值(LOD)由QTL ICIMapping作图软件ICM使用1 000次假设检验和0.05的Ⅰ型错误决定的。经检验阈值为3.1。本研究也采用基于混合模型复合区间作图(MCIM)的QTLNetwork 2.1软件[34]对联合环境的表型数据来检测上位性互作、QTL与环境的互作、QTL与密度互作(由于密度处理类似于环境,故将2个密度视为2个环境)。
1.6 QTL片段在关联群体的分布与衍生分析
利用Yang等[35]和Liu等[36]公布的540份自交系的关联群体的基因型和杂优类群对本研究中在高密度下检测到的QTL区段内的标记进行有利等位基因来源(与掖478或R08基因型相同的标记被视为来源相同)和片段相似成分分析。具体数据分析参考易强[37]的方法。对于部分掖478相关系基于系谱关系进行QTL衍生和传递分析。
2 结果与分析
2.1 RIL群体的茎粗表现、联合方差、遗传力和相关分析
RIL群体的双亲掖478和R08的茎粗在同一密度下均未见显著差异,但R08的茎粗随着密度增加显著变细(P<0.05,图1-A)。这与稀植的R08对密度敏感的预期相符。高低密度下RIL群体的茎粗呈现出数量性状的正态分布特征(图1-B)。与R08响应密度趋势类似,较低密度条件,高密度下RIL群体的茎粗变小,且其变异范围缩小(高密度:13.4~16.1 mm vs 低密度:13.8~17.8 mm)。联合方差分析结果显示,所有因子包括基因型、基因型与环境、基因型与密度等均呈现极显著水平(P<0.01,表1)。此外,高密度下茎粗遗传力(48.01%)相较于低密度下(65.03%)有所降低。相关分析显示,2种密度间RIL群体茎粗呈现出中等显著相关(r=0.68***,P<0.000 1)。但不同密度下RIL群体茎粗与大多数农艺性状显著相关(P<0.05,表2)。如高低密度下茎粗与株高、穗位高、单穗穗质量呈极显著正相关(0.29>r≥0.13,P<0.01),而与空秆率、秃尖长相关性不显著。这表明玉米耐密性是一个综合性状。
表1 2个环境2种密度下RIL群体茎粗的联合方差分析(平方和)
Tab.1 Combined analysis of variance for stem diameter in the RIL population under two densities across two environments
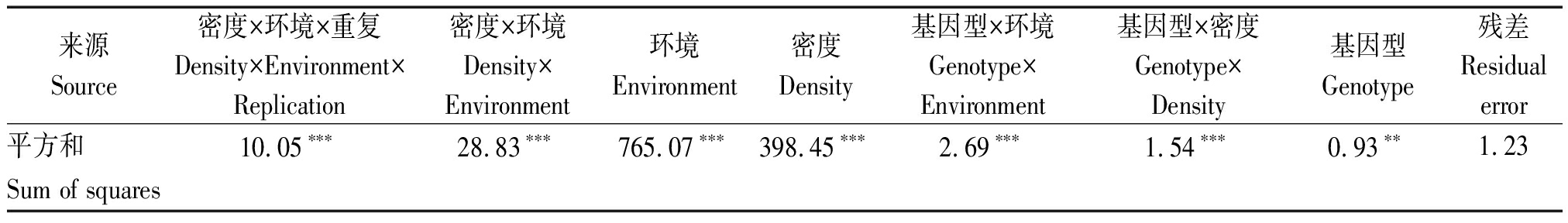
来源Source密度×环境×重复Density×Environment×Replication密度×环境Density×Environment环境Environment密度Density基因型×环境Genotype×Environment基因型×密度Genotype×Density基因型Genotype残差Residualerror平方和10.05∗∗∗28.83∗∗∗765.07∗∗∗398.45∗∗∗2.69∗∗∗1.54∗∗∗0.93∗∗1.23Sum of squares
注:***和**为分别表示P<0.000 1和P<0.01。
Note:*** and ** indicate P<0.000 1 and P<0.01.
表2 2种密度下玉米RIL群体茎粗与其他农艺性状的皮尔森相关系数
Tab.2 Pearson′s correlation coefficients between stem diameter and other agronomic
traits under low and high planting densities in a maize RIL population
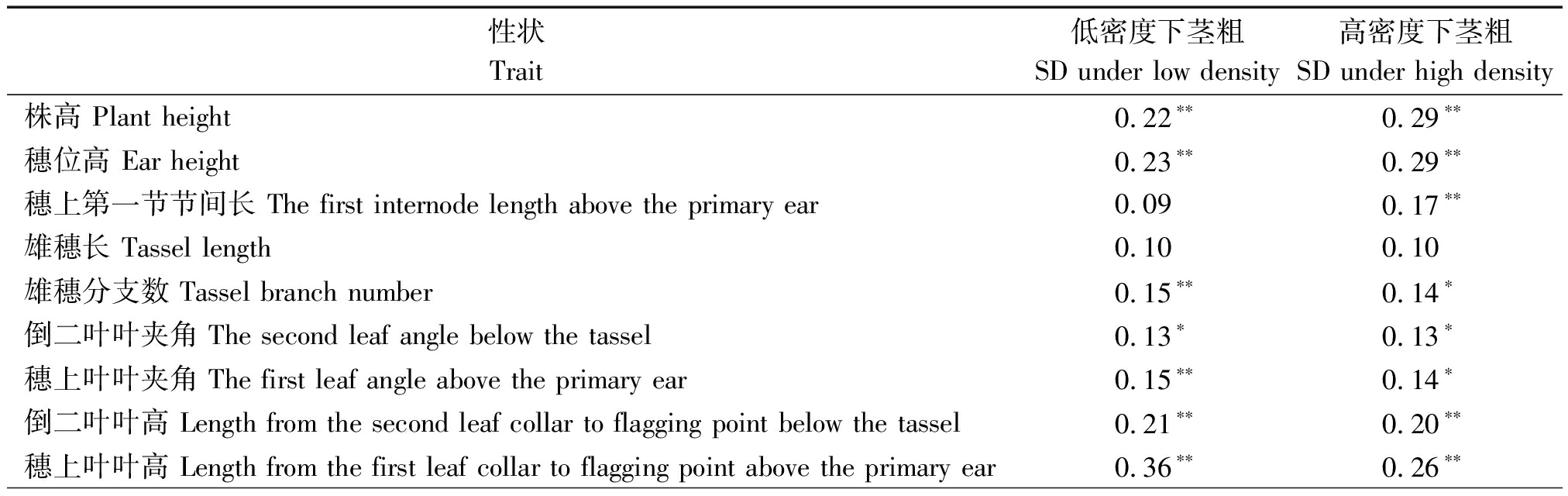

性状Trait低密度下茎粗SD under low density高密度下茎粗SD under high density株高 Plant height0.22∗∗0.29∗∗穗位高 Ear height0.23∗∗0.29∗∗穗上第一节节间长 The first internode length above the primary ear0.090.17∗∗雄穗长 Tassel length0.100.10雄穗分支数 Tassel branch number0.15∗∗0.14∗倒二叶叶夹角 The second leaf angle below the tassel0.13∗0.13∗穗上叶叶夹角 The first leaf angle above the primary ear0.15∗∗0.14∗倒二叶叶高 Length from the second leaf collar to flagging point below the tassel0.21∗∗0.20∗∗穗上叶叶高 Length from the first leaf collar to flagging point above the primary ear0.36∗∗0.26∗∗
2.2 高低密度下RIL群体茎粗的QTL定位分析
本研究对高低密度下RIL群体茎粗分别进行联合环境QTL定位分析,总计发现7个分布于1,3,4,6,7号染色体上的QTL(表3)。这些QTL的表型变异从3.81%~6.91%。其中,qSD1-1、qSD3-1和qSD4仅在高密度种植条件下被检测到,证实这3个QTL均耐高种植密度。位于bin6.05上解释表型变异达6%以上的qSD6同时在2种密度下被观测到,且其增加茎粗的等位基因来源于R08。QTL与环境以及QTL与密度互作的结果未发现存在显著的互作。此外,仅有1对位于1号和10号染色体上的位点显示出加性与加性互作,且其贡献率仅为0.89%(表4)。这些结果显示,在高低密度下茎粗主要受到微效位点控制,而加性和加性互作等效应扮演着更次要的作用。
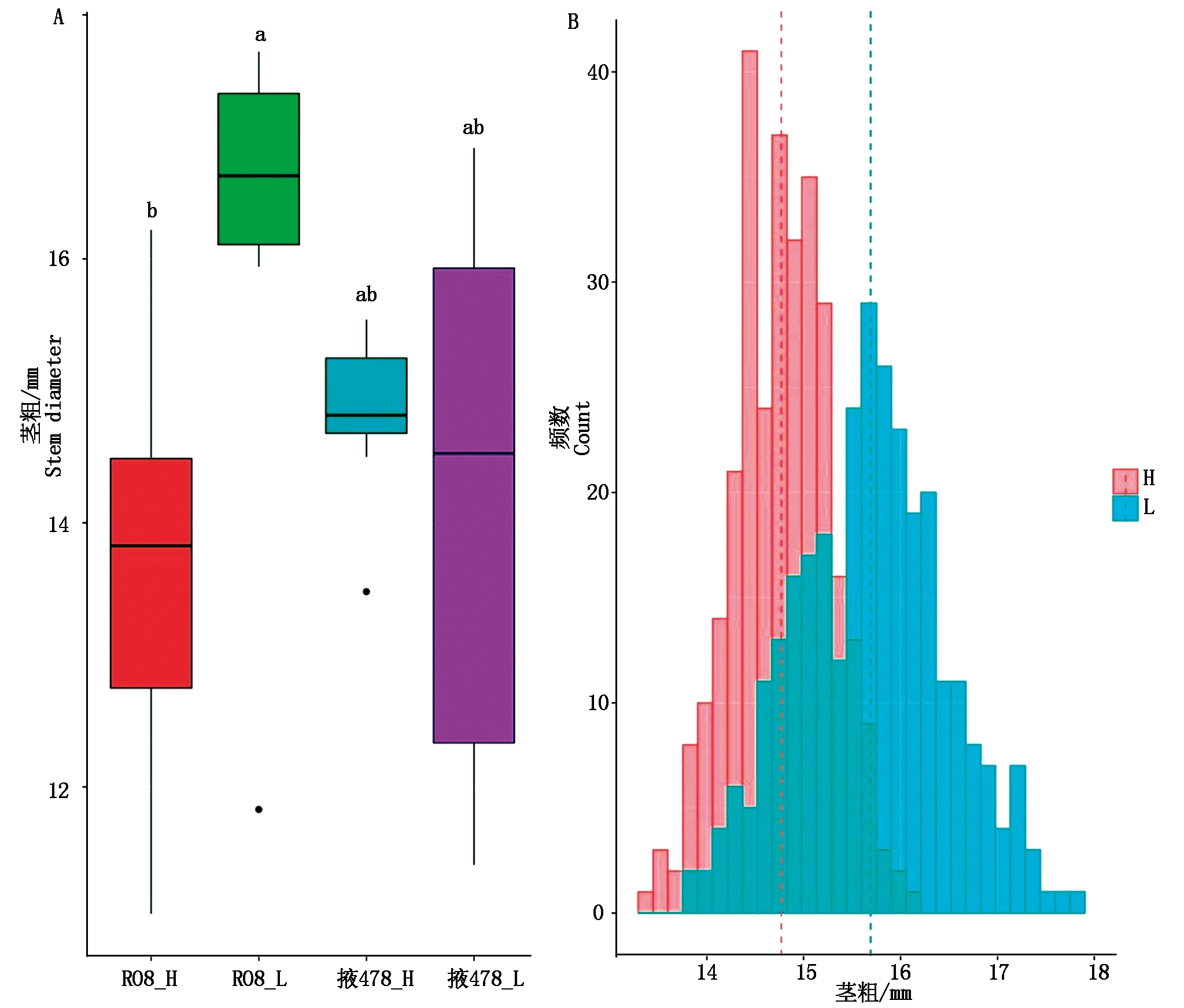
不同小写字母的均值表示存在差异显著(P<0.05)。
Means with different lowercase letters are significant differences at the 0.05 probability level.
图1 掖478、R08(A)及其RIL群体(B)在低(L)和高密度(H)下的茎粗表现
Fig.1 Stem diameter performance for Ye 478,R08(A),and their RIL population(B)under low and high densities
表2(续)
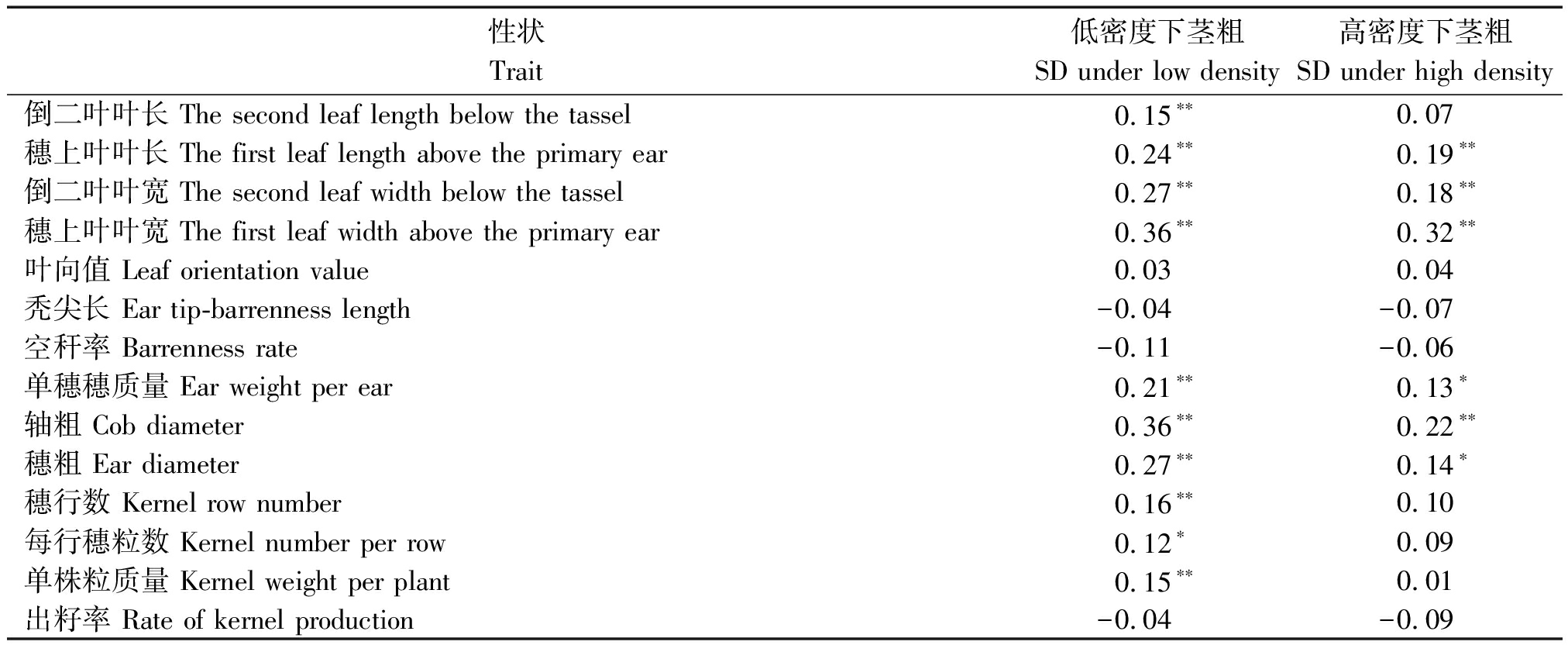

性状Trait低密度下茎粗SD under low density高密度下茎粗SD under high density倒二叶叶长 The second leaf length below the tassel0.15∗∗0.07穗上叶叶长 The first leaf length above the primary ear0.24∗∗0.19∗∗倒二叶叶宽 The second leaf width below the tassel0.27∗∗0.18∗∗穗上叶叶宽 The first leaf width above the primary ear0.36∗∗0.32∗∗叶向值 Leaf orientation value0.030.04秃尖长 Ear tip-barrenness length-0.04-0.07空秆率 Barrenness rate-0.11-0.06单穗穗质量 Ear weight per ear0.21∗∗0.13∗轴粗 Cob diameter0.36∗∗0.22∗∗穗粗 Ear diameter0.27∗∗0.14∗穗行数 Kernel row number0.16∗∗0.10每行穗粒数 Kernel number per row0.12∗0.09单株粒质量 Kernel weight per plant0.15∗∗0.01出籽率 Rate of kernel production-0.04-0.09
注:**和*表示在 P<0.01 和 P<0.05 时显著。
Note:** and * represent significant difference at P<0.01 and P<0.05, respectively.
表3 高低密度下RIL群体茎粗的QTL联合环境定位分析
Tab.3 QTL for stem diameter in the RIL population under low and high densities via joint analysis across two environments
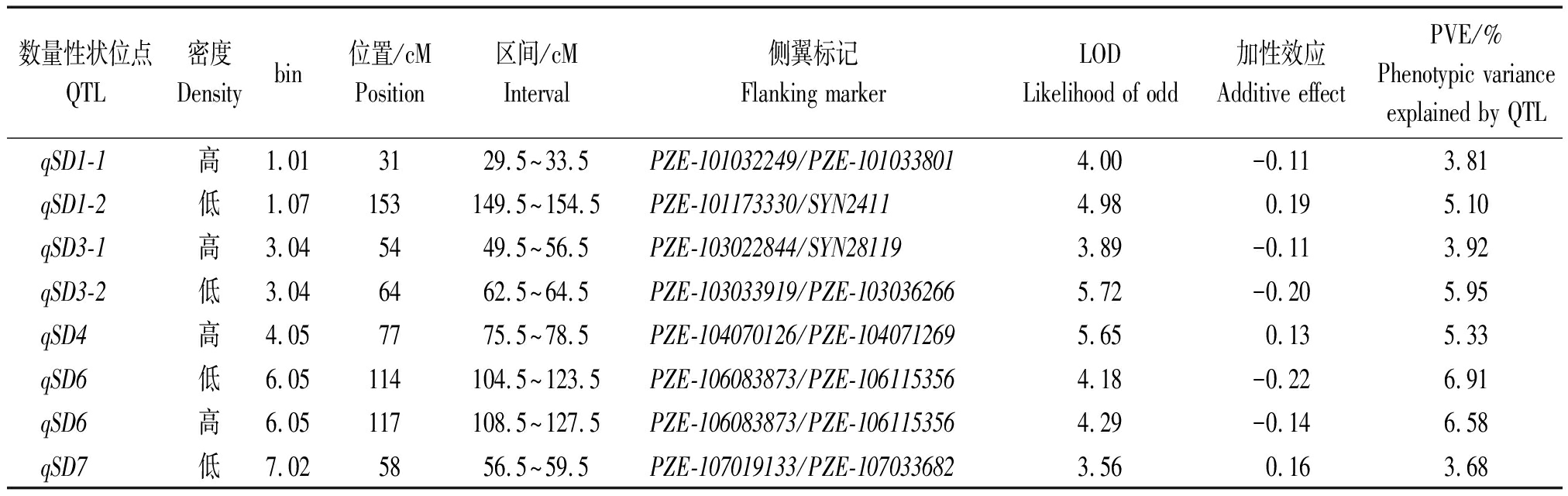
数量性状位点QTL密度Densitybin位置/cMPosition区间/cMInterval侧翼标记Flanking markerLODLikelihood of odd加性效应Additive effectPVE/%Phenotypic varianceexplained by QTLqSD1-1高1.013129.5~33.5PZE-101032249/PZE-1010338014.00-0.113.81qSD1-2低1.07153149.5~154.5PZE-101173330/SYN24114.980.195.10qSD3-1高3.045449.5~56.5PZE-103022844/SYN281193.89-0.113.92qSD3-2低3.046462.5~64.5PZE-103033919/PZE-1030362665.72-0.205.95qSD4高4.057775.5~78.5PZE-104070126/PZE-1040712695.650.135.33qSD6低6.05114104.5~123.5PZE-106083873/PZE-1061153564.18-0.226.91qSD6高6.05117108.5~127.5PZE-106083873/PZE-1061153564.29-0.146.58qSD7低7.025856.5~59.5PZE-107019133/PZE-1070336823.560.163.68
注:QTL命名.q+SD(茎粗)+染色体+系列编号;bin.https://www.maizegdb.org/bin_viewer;PVE.每个QTL解释的表型变异。
Note:QTL nomination.q+SD(Stem diameter)+chromosome+the physical order;bin.https://www.maizegdb.org/bin_viewer;PVE.Phenotypic variance explained by QTL.
表4 高密度下RIL群体茎粗上位性互作分析
Tab.4 Analysis of epistatic interaction for SD in the RIL population under high planting density
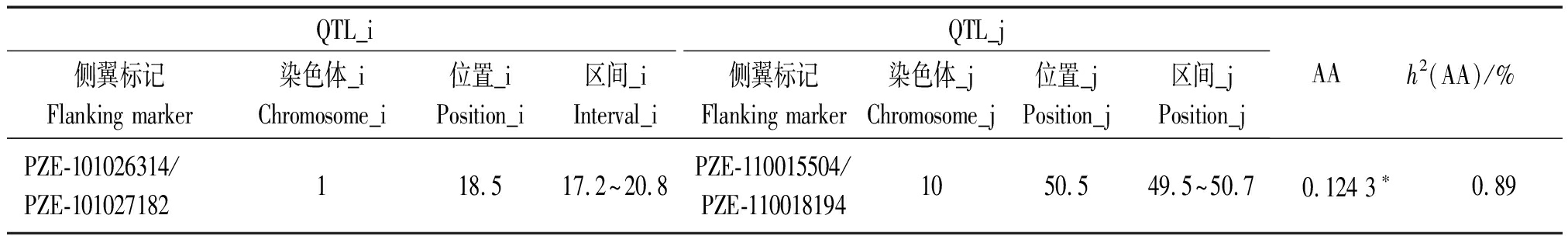
QTL_iQTL_j侧翼标记Flanking marker染色体_iChromosome_i位置_iPosition_i区间_iInterval_i侧翼标记Flanking marker染色体_jChromosome_j位置_jPosition_j区间_jPosition_jAAh2(AA)/%PZE-101026314/PZE-101027182118.517.2~20.8PZE-110015504/PZE-1100181941050.549.5~50.70.124 3∗0.89
注:位置和区间的单位为cM;AA.加性与加性互作;*.P<0.05;h2(AA)为AA的贡献率。
Note:The unit of position and interval is cM;AA.Additive×Additive interaction;*. P<0.05;h2 (AA)is the contribution rate of AA.
2.3 高密度下QTL在关联群体里面的分布和衍生情况
本研究聚焦于高密度下鉴定的qSD1-1、qSD3-1、qSD4和qSD6。除qSD4外,其余3个QTL的增效等位基因均来源于R08。因此,选取这4个QTL观测其在前人报道的关联群体[35-36]的分布情况(图2)。从结果来看,掖478所在的混群(Mix)和R08所在非坚秆群(NSS)在含有利等位基因的QTL区段比例不存在差异,这显示这些区段可能未受到选择。但坚秆群(SS)在这些区段上可能受到一定选择。本研究检测到在7个QTL区段中至少3个QTL区段内来源于掖478的片段衍生或传递率超过70%的自交系达到12个(表5)。除4019、BEM、XUN971和黄C外,其余自交系均为掖478相关系。如从U8112和沈5003聚衍至掖478,再传递至其他相关系。这进一步强调骨干自交系掖478重要区段衍生传递保持自身茎粗相关QTL的特点。
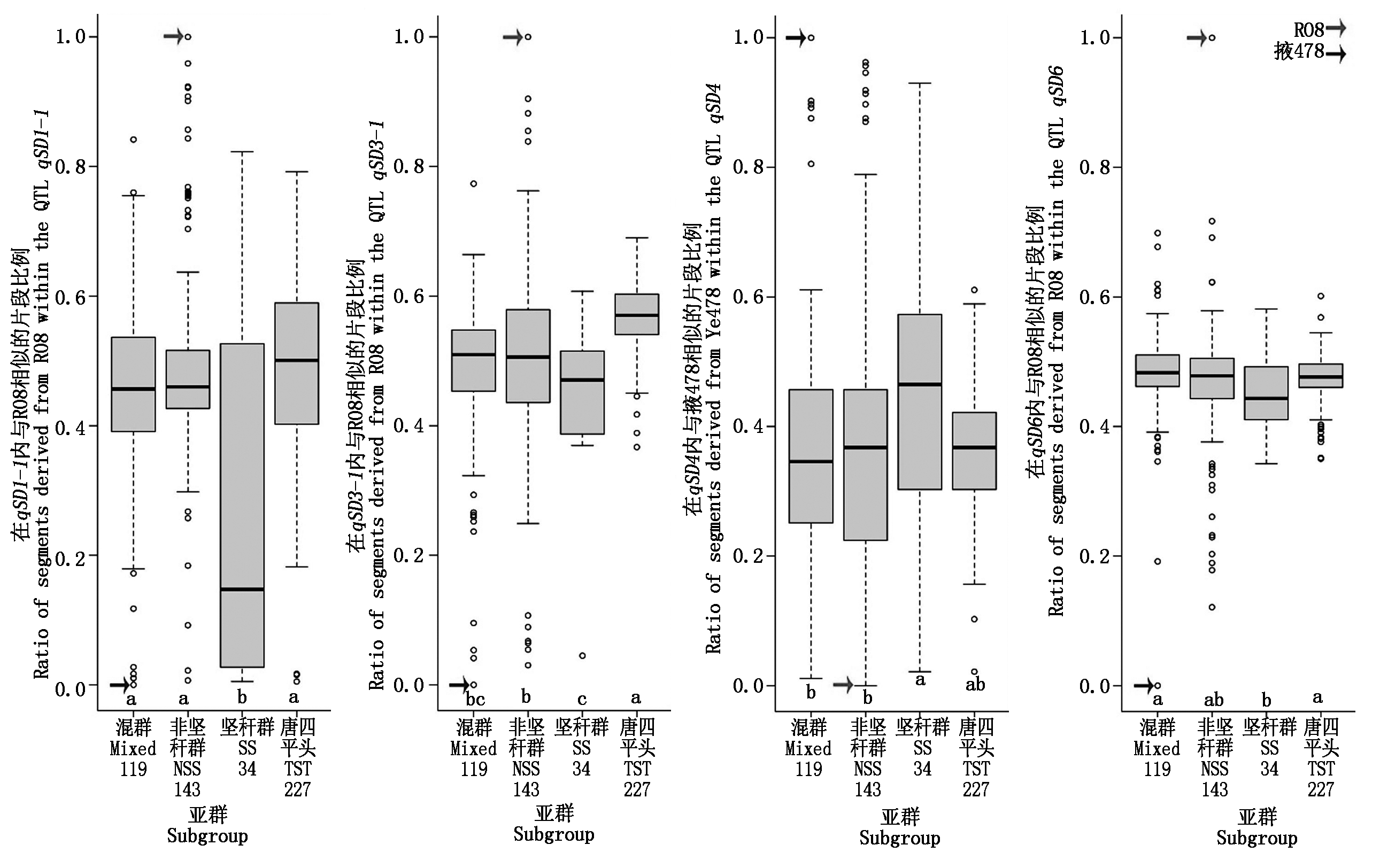
不同字母表示存在显著差异(P<0.05)。
The average with the different letter indicates significant difference(P<0.05).
图2 4个在高种植密度RIL群体下检测到的QTL片段内来源于R08或掖478片段比例在关联群体[35-36]中的分布
Fig.2 Distribution of ratio of segments derived from R08 or Ye478 within four QTL detected in the RIL
population under high planting density across a maize association panel[35-36]
表5 茎粗相关QTL在掖478相关系间的传递率
Tab.5 Transmission rate of QTL for stem diameter among Ye478-related lines
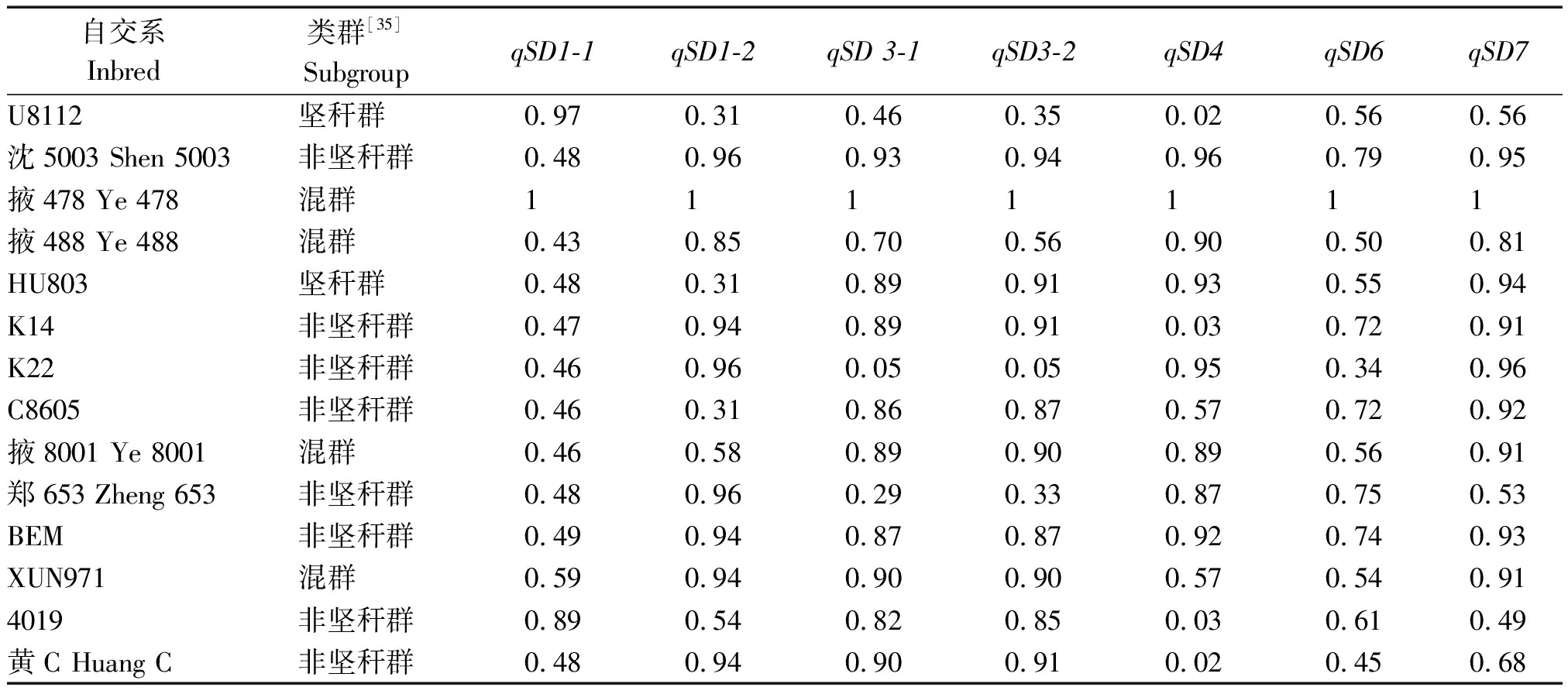
自交系Inbred类群[35]SubgroupqSD1-1qSD1-2qSD 3-1qSD3-2qSD4qSD6qSD7U8112坚秆群0.970.310.460.350.020.560.56 沈5003 Shen 5003非坚秆群0.480.960.930.940.960.790.95 掖478 Ye 478混群1111111 掖488 Ye 488混群0.430.850.700.560.900.500.81 HU803坚秆群0.480.310.890.910.930.550.94 K14非坚秆群0.470.940.890.910.030.720.91 K22非坚秆群0.460.960.050.050.950.340.96 C8605非坚秆群0.460.310.860.870.570.720.92 掖8001 Ye 8001混群0.460.580.890.900.890.560.91 郑653 Zheng 653非坚秆群0.480.960.290.330.870.750.53 BEM非坚秆群0.490.940.870.870.920.740.93 XUN971混群0.590.940.900.900.570.540.91 4019非坚秆群0.890.540.820.850.030.610.49 黄C Huang C非坚秆群0.480.940.900.910.020.450.68
注:区段传递率或衍生率=(每个QTL区段内与掖478分型相同的SNP的数目-每个QTL内R08和掖478相同分型的SNP数目)/每个QTL区段内R08和掖478存在差异的SNP数目。
Note:Segment transmission rate =(Number of SNPs with the same genotypes as Ye478 within each QTL-Number of SNPs with the same genotypes between as Ye 478 and R08 within each QTL)/(Number of SNPs with different genotypes between Ye 478 and R08 within each QTL).
3 结论与讨论
本研究以高低密度下RIL群体茎粗为对象挖掘影响玉米茎粗的位点。研究表明,双亲和RIL的茎粗变异缩小及其遗传力下降的趋势符合预期,与早前报道的其他性状[19-24]相类似。随着群体密度增大,个体间光、热、水和肥竞争压力也随之增大[38-39]。2种密度下RIL群体的茎粗与株高、穗位高和单穗穗质量等极显著相关,而与空秆率和秃尖长等不显著,这与刘鹏飞[10]报道类似。尽管茎粗可能不是评价玉米耐密性的核心指标,但近来研究[11,40-43]显示,玉米茎秆强度和茎粗等与玉米抗倒伏和抗虫害等相关。因此,解析玉米茎秆性状遗传基础对玉米抗逆性仍具有重要意义。
本研究挖掘到7个分布于bin1.01、1.07、3.04(2个)、4.05、6.05和7.02上的控制茎粗的QTL,其中部分QTL在前人研究中也有报道。由于刘福鹏等[12]已收集和整理来自17个不同作图群体的玉米茎粗QTL信息并进行了元分析,因此,本研究未再做Meta分析。本研究低密度下在bin7.02上qSD7与上述研究[12]报道的MQTL13相重合。本研究结果中在2种密度均被观测到的qSD6与徐蔚[6]利用相同来源R08×掖478的F2∶3群体在2个环境下检测到的位点定位到同一区域,但多数QTL未发现有重叠。这证实了家系纯合度、标记数目、定位群体大小和代数及环境数等对QTL定位会产生影响[44-45]。此外,在低密度下位于bin1.07的qSD1-2附近,Zhang等[41]基于关联群体检测到一个与茎粗显著相关的PZE-101172517。在该bin上,刘鹏飞[10]用四交群体F2∶3家系报道了在一个单环境高密度下控制茎粗的qSD-E2-1。尽管众多研究报道了低密度下大量茎粗QTL,但对于高密度下茎秆性状相关报道较少。本研究仅发现一个密度不敏感的qSD6,预示着高低密度下茎粗存在不同遗传调节。因此,提高高种植密度条件下玉米茎秆抗倒伏能力,未来仍需对高种植密度胁迫下遗传来源广泛的玉米自然群体的茎秆性状进行遗传解析。此外,本研究未检测到显著性的QTL与环境以及QTL与密度互作位点,这可能与茎粗遗传力低(相同来源的F2∶3群体中仅略大于45%[6];高低种植密度下RIL群体中分别为48.01%和65.03%)和受环境影响大有关。刘鹏飞[10]仅鉴定到一个环境稳定QTL,而徐蔚[6]在4个环境下未能发现一致性茎粗QTL。本研究显示,茎粗主要受多个微效单位点控制,同时部分受加性与加性互作效应影响,这与前人研究[7,10]略有不同;故全基因分析和选择可能是较为有效选择茎粗的分子辅助手段。
结合笔者之前的报道[19-20],部分影响茎粗的位点也位于多效性QTL区域。在高种植密度下,qSD3-1影响籽粒宽和穗行数;在低密度下,qSD3-2控制容重和穗行数[20]。在bin4.05上,qSD3-2被囊括在影响雄穗分支(高低种植密度)、穗上叶高(高种植密度)和穗上叶长(低种植密度)的多效性区段上[19]。位于bin6.05上2种密度下都检测到的qSD6影响穗上叶长(低种植密度)、节间长(高种植密度)、雄穗分支数(高种植密度)、出籽率(低种植密度)、单穗籽粒质量(低种植密度)和百粒质量(高低种植密度)[19-20]。低密度特异性qSD7包含在一个影响叶长、叶高、空秆率、出籽率和每行穗粒数的多效性位点中[19-20]。因此,通过选择这些多效性位点可间接实现玉米耐密性遗传改良。
尽管本研究多数QTL在广泛遗传背景下的遗传效应未得到证实,但从片段比例选择角度出发,这些区段并未受到选择。这可能与抗倒伏相关性状长期不是育种家的主要目标性状有关。即便骨干亲本掖478相关系中这些区段较为保守,这很可能是选择其他主要农艺性状一因多效的结果。随着生产中对玉米抗倒伏和抗虫害等能力的重视,针对茎秆强度性状的选育将逐渐成为育种家的目标之一。
[1] 张洪生,赵明,吴沛波,翟延举,姜雯. 种植密度对玉米茎秆和穗部性状的影响[J].玉米科学,2009,17(5):130-133. doi:10.13597/j.cnki.maize.science.2009.05.033.
Zhang H S,Zhao M,Wu P B,Zhai Y J,Jiang W. Effects of the plant density on the characteristics of maize stem and ear[J].Journal of Maize Sciences,2009,17(5):130-133.
[2] 任佰朝,李利利,董树亭,刘鹏,赵斌,杨今胜,王丁波,张吉旺. 种植密度对不同株高夏玉米品种茎秆性状与抗倒伏能力的影响[J].作物学报,2016,42(12):1864-1872. doi:10.3724/SP.J.1006.2016.01864.
Ren B Z,Li L L,Dong S T,Liu P,Zhao B,Yang J S,Wang D B,Zhang J W. Effects of plant density on stem traits and lodging resistance of summer maize hybrids with different plant heights[J].Acta Agronomica Sinica,2016,42(12):1864-1872.
[3] 谷利敏,乔江方,张美微,朱卫红,黄璐,代书桃,董树亭,刘京宝. 种植密度对不同耐密夏玉米品种茎秆性状与抗倒伏能力的影响[J].玉米科学,2017,25(5):91-97. doi:10.13597/j.cnki.maize.science.20170515.
Gu L M,Qiao J F,Zhang M W,Zhu W H,Huang L,Dai S T,Dong S T,Liu J B. Effect of planting density on stalk characteristics and lodging-resistant capacity of different density-resistant summer maize varieties[J].Journal of Maize Sciences,2017,25(5):91-97.
[4] 丰光,景希强,李妍妍,王亮,黄长玲. 玉米茎秆性状与倒伏性的相关和通径分析[J].华北农学报,2010,25(S1):72-74. doi:10.7668/hbnxb.2010.S1.017.
Feng G,Jing X Q,Li Y Y,Wang L,Huang C L. Correlation and path analysis of lodging resistance with maize stem characters[J].Acta Agriculturae Boreali-Sinica,2010,25(S1):72-74.
[5] 王铁固,马娟,张怀胜,陈士林. 玉米茎粗主基因+多基因遗传模型分析[J].江苏农业学报,2012,28(3):467-471.doi:10.3969/j.issn.1000-4440.2012.03.002.
Wang T G,Ma J,Zhang H S,Chen S L. Mixed major gene plus polygene inheritance model for stem diameter in maize[J].Jiangsu Journal of Agricultural Sciences,2012,28(3):467-471.
[6] 徐蔚.玉米茎粗QTL定位与遗传分析[D].雅安:四川农业大学,2014.
Xu W. Corn stem diameter QTL mapping and genetic analysis[D].Ya′an:Sichuan Agricultural University,2014.
[7] Hu H X,Liu W X,Fu Z Y,Homann L,Technow F,Wang H W,Song C L,Li S T,Melchinger A E,Chen S J. QTL mapping of stalk bending strength in a recombinant inbred line maize population[J].Theoretical and Applied Genetics,2013,126(9):2257-2266. doi:10.1007/s00122-013-2132-7.
[8] Lemmon Z H,Doebley J F. Genetic dissection of a genomic region with pleiotropic effects on domestication traits in maize reveals multiple linked QTL[J].Genetics,2014,198(1):345-353. doi:10.1534/genetics.114.165845.
[9] 陈琼. 基于DH家系的玉米植株抗倒伏相关性状的QTL分析[D].郑州:河南农业大学,2016.
Chen Q. QTL analysis for the lodging resistance relevant of DH lines about maize plants[D].Zhengzhou:Henan Agricultural University,2016.
[10] 刘鹏飞. 基于四交群体的玉米耐密性及相关性状QTL定位与分析[D].兰州:甘肃农业大学,2013.
Liu P F. QTL mapping of density tolerance and related traits based on four-way cross population in maize(Zea mays L.)[D].Lanzhou:Gansu Agricultural University,2013.
[11] Mazaheri M,Heckwolf M,Vaillancourt B,Gage J L,Burdo B,Heckwolf S,Barry K,Lipzen A,Ribeiro C B,Kono T J Y,Kaeppler H F,Spalding E P,Hirsch C N,Robin Buell C,de Leon N,Kaeppler S M. Genome-wide association analysis of stalk biomass and anatomical traits in maize[J].BMC Plant Biology,2019,19(1):45. doi:10.1186/s12870-019-1653-x.
[12] 刘福鹏,曲文利,房海悦,李莉莉,金峰学,吴委林. 玉米茎粗Meta-QTL及候选基因分析[J].东北农业科学,2019,44(5):30-33. doi:10.16423/j.cnki.1003-8701.2019.05.007.
Liu F P,Qu W L,Fang H Y,Li L L,Jin F X,Wu W L. Analysis of Meta-QTL and candidate genes related to stem diameter in maize[J].Journal of Northeast Agricultural Sciences,2019,44(5):30-33.
[13] Gonzalo M,Vyn T J,Holland J B,McIntyre L M. Mapping density response in maize:A direct approach for testing genotype and treatment interactions[J].Genetics,2006,173(1):331-348. doi:10.1534/genetics.105.045757.
[14] Gonzalo M,Holland J B,Vyn T J,McIntyre L M. Direct mapping of density response in a population of B73Mo17 recombinant inbred lines of maize(Zea Mays L.)[J].Heredity,2010,104(6):583-599. doi:10.1038/hdy.2009.140.
[15] Ku L X,Zhang L K,Tian Z Q,Guo S L,Su H H,Ren Z Z,Wang Z Y,Li G H,Wang X B,Zhu Y G,Zhou J L,Chen Y H. Dissection of the genetic architecture underlying the plant density response by mapping plant height-related traits in maize(Zea mays L.)[J].Molecular Genetics and Genomics,2015,290(4):1223-1233. doi:10.1007/s00438-014-0987-1.
[16] Ku L X,Ren Z Z,Chen X,Shi Y,Qi J S,Su H H,Wang Z Y,Li G H,Wang X B,Zhu Y G,Zhou J L,Zhang X,Chen Y H. Genetic analysis of leaf morphology underlying the plant density response by QTL mapping in maize(Zea mays L.)[J].Molecular Breeding,2016,36(5):1-16. doi:10.1007/s11032-016-0483-x.
[17] Wang H W,Liang Q J,Li K,Hu X J,Wu Y J,Wang H,Liu Z F,Huang C L. QTL analysis of ear leaf traits in maize(Zea mays L.)under different planting densities[J].The Crop Journal,2017,5(5):387-395. doi:10.1016/j.cj.2017.05.001.
[18] 任真真,周金龙,王小博,李国辉,朱宇光,库丽霞,陈彦惠. 两种密度条件下玉米穗上节间距QTL分析[J].玉米科学,2016,24(3):26-30. doi:10.13597/j.cnki.maize.science.20160305.
Ren Z Z,Zhou J L,Wang X B,Li G H,Zhu Y G,Ku L X,Chen Y H. QTL mapping of internodes length above upmost ear under two planting densities in maize[J].Journal of Maize Sciences,2016,24(3):26-30.
[19] Yi Q,Hou X B,Liu Y H,Zhang X G,Zhang J J,Liu H M,Hu Y F,Yu G W,Li Y P,Huang Y B. QTL analysis for plant architecture-related traits in maize under two different plant density conditions[J].Euphytica,2019,215:148. doi:10.1007/s10681-019-2446-x.
[20] Yi Q,Liu Y H,Hou X B,Zhang X G,Zhang J J,Liu H M,Hu Y F,Yu G W,Li Y P,Wang Y B,Huang Y B. Quantitative trait loci mapping for yield-related traits under low and high planting densities in maize(Zea mays)[J].Plant Breeding,2020,139(2):227-240.doi:10.1111/pbr.12778.
[21] Guo J J,Chen Z L,Liu Z P,Wang B B,Song W B,Li W,Chen J,Dai J R,Lai J S. Identification of genetic factors affecting plant density response through QTL mapping of yield component traits in maize(Zea mays L.)[J].Euphytica,2011,182(3):409-422. doi:10.1007/s10681-011-0517-8.
[22] 王辉,梁前进,胡小娇,李坤,黄长玲,王琪,何文昭,王红武,刘志芳. 不同密度下玉米穗部性状的QTL分析[J].作物学报,2016,42(11):1592-1600. doi:10.3724/SP.J.1006.2016.01592.
Wang H,Liang Q J,Hu X J,Li K,Huang C L,Wang Q,He W Z,Wang H W,Liu Z F. QTL mapping for ear architectural traits under three plant densities in maize[J].Acta Agronomica Sinica,2016,42(11):1592-1600.
[23] 刘翔攀. 玉米自交系耐密性评价及SNP关联分析[D].泰安:山东农业大学,2017.
Liu X P. Evaluation on density-tolerant characteristics of maize inbred lines and association analysis between SNP and density-tolerance[D].Tai′an:Shandong Agricultural University,2017.
[24] Wang L W,Zhou Z Q,Li R G,Weng J F,Zhang Q G,Li X H,Wang B Q,Zhang W Y,Song W,Li X H. Mapping QTL for flowering time-related traits under three plant densities in maize[J].The Crop Journal,2021,9(2):372-379. doi:10.1016/j.cj.2020.07.009.
[25] Littell R C,Milliken G A,Stroup W W,Wolfinger R D,Schabenberger O. SAS for mixed models[M].2nd edition. North Carolina:SAS Institute,Cary,2007.
[26] Holland J B,Nyquist W E,Cervantes-Martínez C T. Estimating and interpreting heritability for plant breeding:An update[M]//Plant Breeding Reviews. Oxford:John Wiley & Sons,Inc.,2010:9-112. doi:10.1002/9780470650202.ch2.
[27] The R Development Core Team. R:A language and environment for statistical computing[EB/OL].R Foundation for Statistical Computing,Vienna,Austria,2015.
[28] Bates D,Mächler M,Bolker B,Walker S. Fitting linear mixed-effects models Usinglme 4[J].Journal of Statistical Software,2015,67(1):137528.doi:10.18637/jss.v067.i01.
[29] Chen D H,Ronald P C. A rapid DNA minipreparation method suitable for AFLP and other PCR applications[J].Plant Molecular Biology Reporter,1999,17(1):53-57. doi:10.1023/A:1007585532036.
[30] Ganal M W,Durstewitz G,Polley A,Bérard A,Buckler E S,Charcosset A,Clarke J D,Graner E M,Hansen M,Joets J,Le Paslier M C,McMullen M D,Montalent P,Rose M,Schön C C,Sun Q,Walter H,Martin O C,Falque M. A large maize(Zea mays L.)SNP genotyping array:Development and germplasm genotyping,and genetic mapping to compare with the B73 reference genome[J].PLoS One,2011,6(12):e28334. doi:10.1371/journal.pone.0028334.
[31] Lorieux M. MapDisto:fast and efficient computation of genetic linkage maps[J].Molecular Breeding,2012,30(2):1231-1235. doi:10.1007/s11032-012-9706-y.
[32] Kosambi D D. The estimation of map distances from recombination values[J].Annals of Eugenics,1943,12(1):172-175.doi:10.1111/j.1469.1809.1943.tb02321.x.
[33] Meng L,Li H H,Zhang L Y,Wang J K. QTL IciMapping:Integrated software for genetic linkage map construction and quantitative trait locus mapping in biparental populations[J].The Crop Journal,2015,3(3):269-283. doi:10.1016/j.cj.2015.01.001.
[34] Yang J,Hu C C,Hu H,Yu R D,Xia Z,Ye X Z,Zhu J. QTLNetwork:mapping and visualizing genetic architecture of complex traits in experimental populations[J].Bioinformatics,2008,24(5):721-723. doi:10.1093/bioinformatics/btm494.
[35] Yang X H,Gao S B,Xu S T,Zhang Z X,Prasanna B M,Li L,Li J S,Yan J B. Characterization of a global germplasm collection and its potential utilization for analysis of complex quantitative traits in maize[J].Molecular Breeding,2011,28(4):511-526. doi:10.1007/s11032-010-9500-7.
[36] Liu H J,Luo X,Niu L Y,Xiao Y J,Chen L,Liu J,Wang X Q,Jin M L,Li W Q,Zhang Q H,Yan J B. Distant eQTLs and non-coding sequences play critical roles in regulating gene expression and quantitative trait variation in maize[J].Molecular Plant,2017,10(3):414-426. doi:10.1016/j.molp.2016.06.016.
[37] 易强. 玉米骨干亲本掖478和08-641关键区段的遗传解析[D].成都:四川农业大学,2020.
Yi Q. Genetic dissection of specific genomic regions for maize foundation parents Ye478 and 08-641[D].Chengdu:Sichuan Agricultural University,2020.
[38] Tetio-Kagho F,Gardner F P. Responses of maize to plant population density.Ⅰ.Canopy development,light relationships,and vegetative growth[J].Agronomy Journal,1988,80(6):930-935. doi:10.2134/agronj1988.00021962008000060018x.
[39] Tetio-Kagho F,Gardner F P. Responses of maize to plant population density.Ⅱ.Reproductive development,yield,and yield adjustments[J].Agronomy Journal,1988,80(6):935-940.doi:10.2134/agronj1988.00021962008000060019x.
[40] Liu X G,Hu X J,Li K,Liu Z F,Wu Y J,Wang H W,Huang C L. Genetic mapping and genomic selection for maize stalk strength[J].BMC Plant Biology,2020,20(1):196. doi:10.1186/s12870-020-2270-4.
[41] Zhang Y L,Liu P,Zhang X X,Zheng Q,Chen M,Ge F,Li Z L,Sun W T,Guan Z R,Liang T H,Zheng Y,Tan X L,Zou C Y,Peng H W,Pan G T,Shen Y O. Multi-locus genome-wide association study reveals the genetic architecture of stalk lodging resistance-related traits in maize[J].Frontiers in Plant Science,2018,9:611. doi:10.3389/fpls.2018.00611.
[42] Zhang Y,Wang J L,Du J J,Zhao Y X,Lu X J,Wen W L,Gu S H,Fan J C,Wang C,Wu S,Wang Y J,Liao S J,Zhao C J,Guo X Y. Dissecting the phenotypic components and genetic architecture of maize stem vascular bundles using high-throughput phenotypic analysis[J].Plant Biotechnology Journal,2021,19(1):35-50. doi:10.1111/pbi.13437.
[43] López-Malvar A,Butrón A,Samayoa L F,Figueroa-Garrido D J,Malvar R A,Santiago R. Genome-wide association analysis for maize stem cell wall-bound hydroxycinnamates[J].BMC Plant Biology,2019,19(1):519. doi:10.1186/s12870-019-2135-x.
[44] Yi Q,Liu Y H,Zhang X G,Hou X B,Zhang J J,Liu H M,Hu Y F,Yu G W,Huang Y B. Comparative mapping of quantitative trait loci for tassel-related traits of maize in F2∶3 and RIL populations[J].Journal of Genetics,2018,97(1):253-266. doi:10.1007/s12041-018-0908-x.
[45] Yi Q,Liu Y H,Hou X B,Zhang X G,Li H,Zhang J J,Liu H M,Hu Y F,Yu G W,Li Y P,Wang Y B,Huang Y B. Genetic dissection of yield-related traits and mid-parent heterosis for those traits in maize(Zea mays L.)[J].BMC Plant Biology,2019,19(1):392. doi:10.1186/s12870-019-2009-2.